Access Chipster
Access Chipster
Introduction
This guide is intended for researchers who want to use Chipster (v3.12.3), an user-friendly software for analyzing high-throughput data such as NGS and microarrays, in the cloud-based resources provided by EGI.
The software contains over 400 analysis tools and a large collection of reference genomes.
Users can save and share automatic analysis workflows, and visualize data interactively using for example the built-in genome browser.
The Chipster testbed in EGI resources
The Chipster testbed configured at CESGA offers:
- 8 vCPU cores,
- 32GB of RAM,
- 1TB of block storage in /data,
- Software and tools (v3.12.3) are in available under the /cvmfs/chipster.egi.eu/ partition.
This how-to has been tested in the EGI Applications on Demand Infrastructure.
The only pre-requisite to access and use the Chipster testbed is to be a member of this infrastructure.
About this wiki
In this guide we will show how to:
- Create a temporary Chipster account;
- Use the temporary account to access the testbed and run the bio-informatics applications.
How to create/review a temporary account
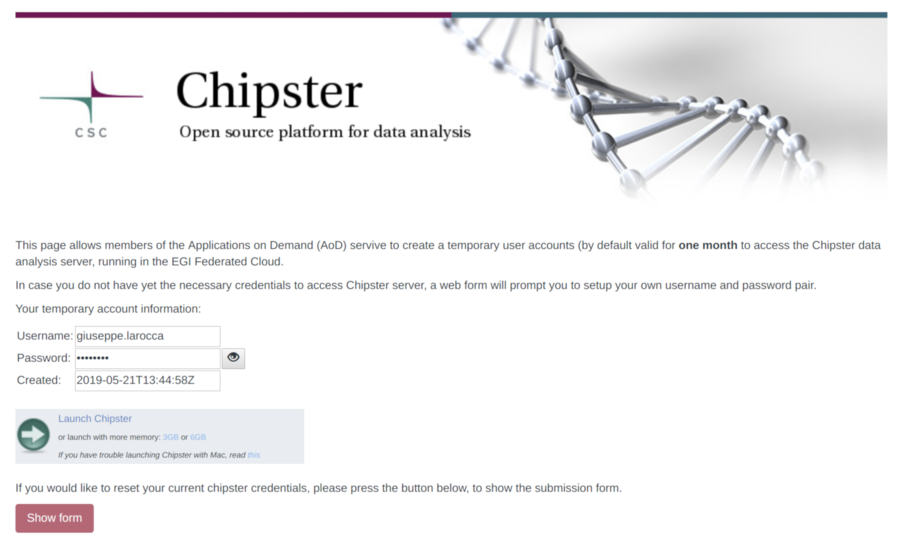
In this section we will show how to create/renew your temporary Chipster account. By default, a temporary account to access the Chipster server is valid for one month.
To generate/review your temporary Chipster account, please use this link.
- Click on the "Show form".
- If the account has expired, the Science Gateway will automatically generate a new password for the user.
- To activate the new account, click on the "Execute" button to trigger the creation/update of the temporary account in the Chipster testbed. This operation may takes few minutes.
- User can monitor the execution of the task clicking in the "Show" button.
- When the account is create/renewed, the web interface shows a link to access the Chipster server with the new credentials.
Acknowledgement
Users of the service are asked to provide appropriate acknowledgement of the use in scientific publications. The following acknowledgement text can be used for this purpose: This work used the EGI Applications on Demand service, which is co-funded by the EOSC-hub project (grant number 777536).