Difference between revisions of "Access Chipster"
| Line 2: | Line 2: | ||
== Introduction == | == Introduction == | ||
This guide is intended for researchers who want to use [http://chipster.csc.fi/ Chipster (v3.12.3)], an user-friendly software for analyzing high-throughput data such as NGS and microarrays, in the cloud-based resources provided by [https://wiki.egi.eu/wiki/Long-tail_of_science EGI]. | This guide is intended for researchers who want to use [http://chipster.csc.fi/ Chipster (v3.12.3)], an user-friendly software for analyzing high-throughput data such as NGS and microarrays, in the cloud-based resources provided by [https://wiki.egi.eu/wiki/Long-tail_of_science EGI]. The software contains over 400 analysis tools and a large collection of reference genomes. Users can save and share automatic analysis workflows, and visualize data interactively using for example the built-in genome browser. | ||
The Chipster server configured at [https://appdb.egi.eu/store/site/cesga CESGA] offers: | The Chipster server configured at [https://appdb.egi.eu/store/site/cesga CESGA] offers: | ||
Revision as of 11:23, 3 June 2019
Access Chipster
Introduction
This guide is intended for researchers who want to use Chipster (v3.12.3), an user-friendly software for analyzing high-throughput data such as NGS and microarrays, in the cloud-based resources provided by EGI. The software contains over 400 analysis tools and a large collection of reference genomes. Users can save and share automatic analysis workflows, and visualize data interactively using for example the built-in genome browser.
The Chipster server configured at CESGA offers:
- 8 vCPU cores,
- 32GB of RAM,
- 1TB in /data
- Software and tools (v3.12.3) are in /cvmfs/chipster.egi.eu/
This how-to has been tested in the EGI Applications on Demand Infrastructure.
The only pre-requisite to access and use the Chipster testbed is to be a member of this infrastructure.
Objectives
In this guide we will show how to
- Create a temporary Chipster account to access the testbed;
- Use the temporary account to access and run the bio-informatics applications.
How to access the Chipster testbed
To access the Chipster tested, a valid account is needed.
- If the user already HAS a valid user name and password for this Chipster service, he/she can launch the Chipster client from the Chipster
- If the user did NOT use this Chipster server before or if username and password have expired, please use this link to generate a new one (see next section for more details).
- By default, the password will be valid for one month.
How to create a temporary account
In this section we will show how to create/renew your temporary account using the JSR 286 portlet available in the FutureGateway Science Gateway.
By default, a temporary account to access the Chipster server is valid for one month.
To create/renew your temporary account:
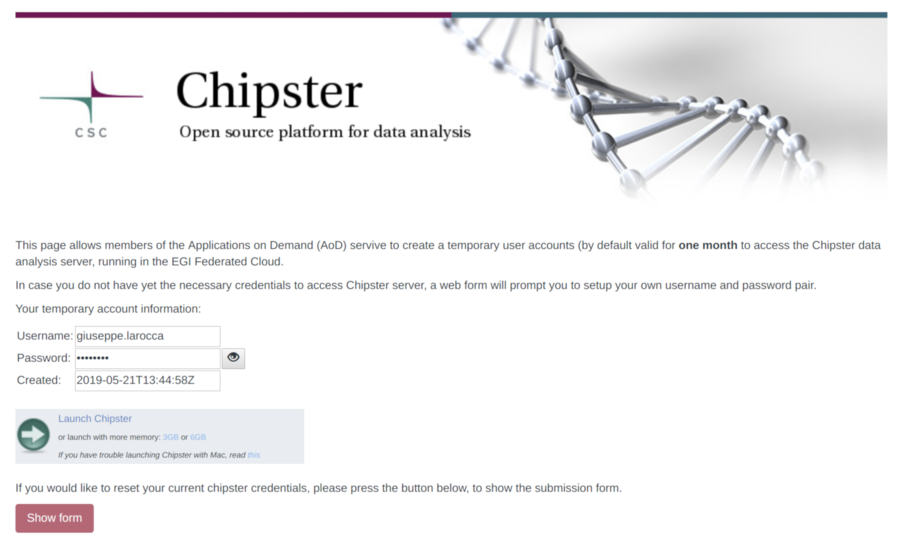
- Click on the "Show form" button as showed below. By default, the Science Gateway will automatically generate a new password for the user.
- Click on "Execute" to trigger the creation/update of the account in the Chipster testbed. You can monitor the execution of the task clicking in the "Show" button.
- When the account is create/renewed, the user can access the server